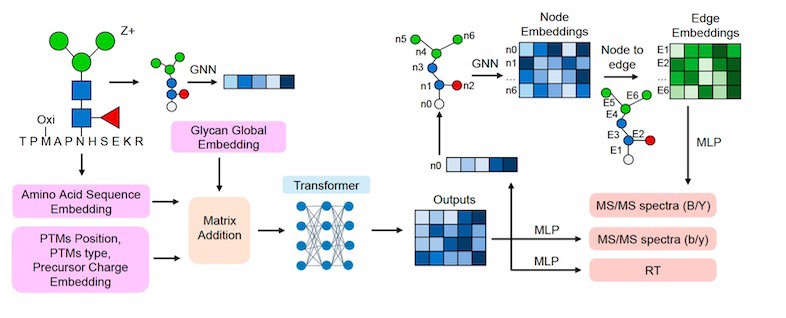
Protein glycosylation, a post-translational modification of proteins by glycans, plays an important role in numerous physiological and pathological cellular functions. Glycoproteomics, the study of protein glycosylation on a proteome-wide scale, utilizes liquid chromatography coupled with tandem mass spectrometry (MS/MS) to get combinational information on glycosylation site, glycosylation level and glycan structure. However, current database searching methods for glycoproteomics often struggle with glycan structure determination due to the limited occurrence of structure-determining ions. Although spectral searching methods can leverage fragment intensity to facilitate the structure identification of glycopeptides, their application is hindered by difficulties in spectral library construction. In this work, we present DeepGP, a hybrid deep learning framework based on transformer and graph neural networks, for the prediction of MS/MS spectra and retention time of glycopeptides. Two graph neural network modules are employed to capture the branched glycan structure and predict glycan ion intensity, respectively. Additionally, a pretraining strategy is implemented to alleviate the insufficiency of glycoproteomics data. Testing on multiple biological datasets, DeepGP accurately predicts MS/MS spectra and retention time of glycopeptides, closely aligning with the experimental results. Comprehensive benchmarking of DeepGP on synthetic and biological datasets validates its effectiveness in distinguishing similar glycans. Based on various decoy methods, DeepGP in combination with database searching can increase glycopeptide detection sensitivity. We anticipate that DeepGP can inspire extensive future work in glycoproteomics.
https://www.nature.com/articles/s42256-024-00875-x